Which Is The Best Model Organism?

Science is complicated. From the unimaginable scales of the universe to the numerous interactions between molecules in space, just the physical aspects of science can be practically impossible for us to accurately interpret. Even for veterans and experts in their field of study, much of the time their investigations focus not on the entirety of a subject, but instead on a small section that (hopefully!) fits into the whole. In fact, this is also one of the reasons that mathematics can be so useful sometimes. When researchers require a general representation of a process, depicting it with a mathematical model can be the key to viewing the bigger picture.
In the life sciences, there is no shortage of models. Besides being known for the typical prototypes of DNA, cells and animal anatomies that we often see in schools, they are also famous for their model organisms. These include bacteria like Escherichia coli; unicellular or multicellular fungi, such as the bioluminescent Neonothopanus gardneri; or even the ubiquitous and almighty fruit fly, Drosophila melanogaster. Unlike pedagogical figures, model organisms are used primarily to facilitate research. Some of them are used as tools for genetic replication and engineering (E. coli being one such example), while a few are needed for the internal environments they provide to biological samples that we'd like to observe. Yet others give researchers the opportunity to explore more deeply into metabolic pathways or unique cellular behaviours, discovering esoteric mechanisms that are seen in no other place on the planet. As a biochemist, I am surrounded by these kinds of models. And, though they might not be the most practical thing to learn for the average folk, I think it's a wonderful way to get a little bit of perspective into what researchers actually do.
Evolutionary Yeast
It may interest you to know that the yeast we use for bread-baking is actually a type of fungus. Brewer's yeast, or Saccharomyces cerevisiae, is a unicellular organism that is widely entertained in labs not for its ability to ferment sugar - but for how simple it is to cultivate and modify and its physiology as a eukaryote. This means that, like our own body cells, yeast cells contain membrane-bound organelles that separate different metabolic pathways that could otherwise conflict with each other if they occurred in the same space. Their genome is also surprisingly similar to ours, with 23 percent of functional genes in yeast being homologous to human genes. The genomes are not completely identical, of course - the one in S. cerevisiae contains a bit over 6000 genes, for instance, with human cells having over 20000 of them - but they are still sufficiently related that studying the fungus lets us make good predictions for how our own cells might work. The same is true for many other multicellular eukaryotes (though it does sound rather bizarre to compare yeast with whales and such).
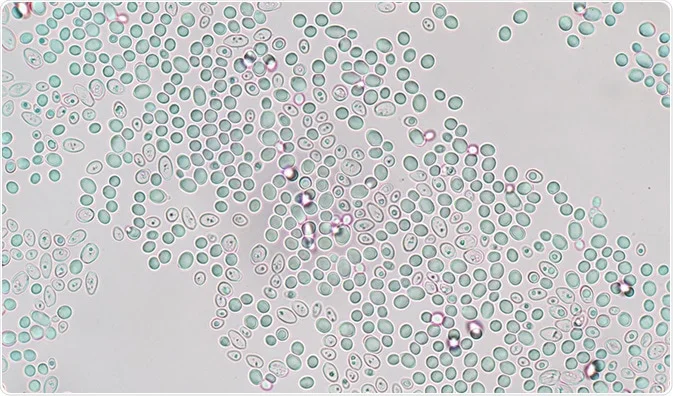
I should note that the difference in sizes between human and yeast genomes is not that reflective of their complexity. In reality, most of that difference comes from genes or non-functional sequences being duplicated in the genetic code over the millennia. Ultimately, this can lead to humans bearing extended DNA sections that (appear to) serve no more purpose than a corresponding smaller genetic sequence in yeast DNA. Occasionally, of course, the duplicated sequence does serve a functional purpose.
Both yeast and humans contain intersectin proteins, which are known to attach to other proteins to facilitate the formation of vesicles that transport external materials into the cell (a widely studied process called endocytosis). In humans, two of these proteins exist - intersectin-1 and intersectin-2, the former of which has the additional role of activating actin polymerisation around the premature vesicle. Yeast intersectin is a different story. Known in S. cerevisiae as Pan-1 and Sla-1, these proteins are thought to be two halves of a larger intersectin that got divided in the past. From this, we can conclude two things: that there existed a common ancestor between humans and yeast that possessed a fully intact intersectin gene; and that diverging evolution led it to be duplicated in humans while it was split in yeast cells.
Whether through natural selection or genetic drift, model organisms can tell us how evolution has changed different species from each other over time. And this is true not only by comparing us with distantly related yeast cells, but even with other primates. Here, it may be appropriate to make something clear - humans do not come from gorillas. We didn't evolve from orangutans, nor from chimpanzees or bonobos. Instead, all primates diverged originally from a common ancestor (although this particular ancestor would have lived more recently than the human-yeast ancestor). It is the job of evolutionary biologists to figure out why and how an organism's DNA got altered, and such is made that much easier by contrasting it with the genomes in other model organisms.

Altogether, we can compare the molecular characteristics of an entire community of organisms to more accurately predict the shapes and DNA of past lifeforms. This is one of the reasons why we don't only use one yeast type - we also use budding yeast (Schizosaccharomyces pombe) and corn smut (Ustilago maydis), the group of which look mostly undistinguishable in lab conditions but display striking differences in their biochemistry. I've even heard talk that brewer's yeast is evolutionarily as far away from budding yeast as humans are from fruit flies. But I digress.
Besides figuring out evolution, we observe model organisms for their potential in treating ailments and dissecting the complex metabolic pathways in their bodies. Personally, this is probably also what interests me the most, particularly with the boundless variety that exists in life.
Modelling Metabolisms
Rats, rabbits and other rodents are very common model organisms in science (hence the term 'lab mouse'). This is because of the notion that the most important discoveries in biochemistry are applicable in mammals like humans, most often medically. While I don't blame the community for thinking this way - we all want to have a better healthcare, right? - it can become slightly frustrating for those researchers who'd like to investigate the quirks behind bats or axolotls or Boop boop fish (which bear the best species name I've found on Twitter). Nevertheless, rodent research has revolutionised our knowledge in multicellular eukaryotes. Associated discoveries range from how cancer develops and stem cells differentiate to the intricacies in our nervous and immune systems. Once upon a time, virologists even genetically modified mice (Mus musculus) with CD155, a human gene responsible for the protein receptor that allows the Poliovirus to infect us. This gave them the capability to study the triggers and symptoms of Polio, ultimately producing a vaccine that has saved millions since the year 1952.

To research how cells die and the causes behind disorders like congenital heart disease, biologists can also utilise the nematode worm (Caenorhabditis elegans). This little worm has been in laboratories since Ellsworth Dougherty noted in 1948 that free-living nematodes like C. elegans could be useful for genetic study. True to his words, genetic analyses have revealed many-a-protein that regulates apoptosis in cells (a form of programmed cell death) and organ development in eukaryotes. This research then reached a peak when Sydney Brenner, Robert Horvitz and John E. Sulton - each with his own worm - won the Nobel Prize in Physiology or Medicine in 2002. I encourage you to look some of this up later, for it really is fascinating stuff.
In a similar way as with C. elegans, we can visualise biological molecules and their interactions with each other in D. melanogaster, the venerable fruit fly. Fruit flies are far more prevalent than worms, however. Alongside brewer's yeast, this species has been domesticated to a fault over the years, reaching such an ease of maintenance and modification that we now have entire companies dedicated to them. And with good incentive, because we use flies for everything; for evolutionary studies and tissue growth to protein regulation and epigenetics, you'll see them grown in practically every major scientific institution out there.
Fruit fly biology is incredibly well understood. Coupled with how clearly different mutations can affect them, it is easy to see why so many award-winning papers have involved them. Take the Minute gene, which can exist in two forms (i.e., two alleles) in fruit flies. When one of the two Minute gene copies in a fly is mutated, the individual will grow up to have shorter hairs and smaller wings than their wild-type counterparts. When both copies are mutated, the fly dies - all in all demonstrating a recurring phenomenon in genetics called haploinsufficiency. Other genes of interest in flies include the eyeless gene (involved in eye development), the cry gene (which expresses cryptochrome-1 to regulate the circadian clock, as with Monarch butterflies) and many others. Generally, fruit fly genes are named according to the observable characteristics (phenotype) that occur when the gene is mutated. In this sense, why do you think their eyeless gene is named this way?
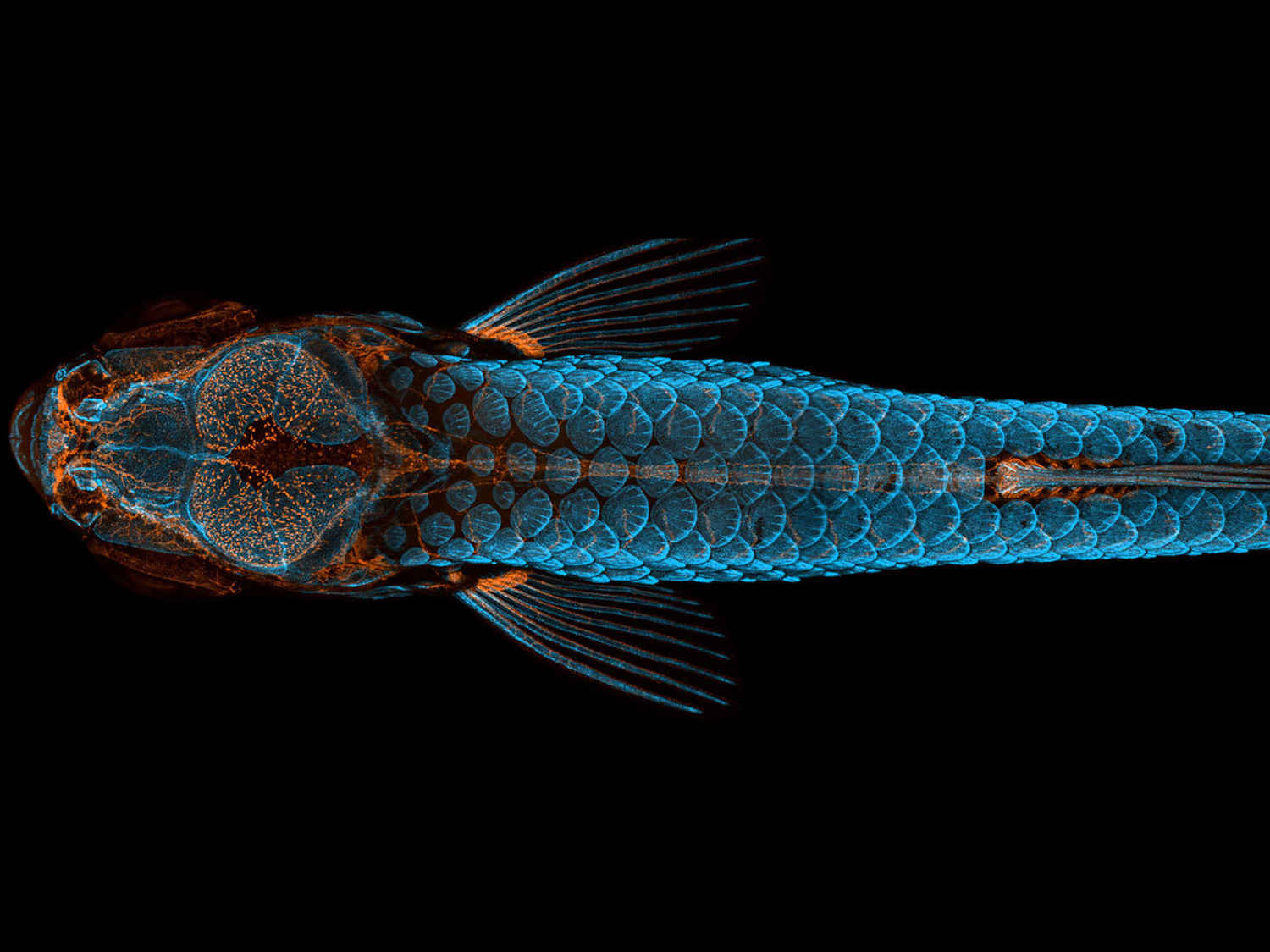
The final common model organism I'd like to talk about is the zebrafish (Danio rerio). Although they require pretty strict conditions to breed, these little minnow grow astoundingly quickly and so are perfect for studying early development. With a nigh-on transparent skin, they are also very undemanding to image with most kinds of microscopy, producing some of the most beautiful fluorescent images in the life sciences. A recent success with them (which I will probably cover in a separate blog post sometime) was the discovery of asynthetic fission by a group of researchers from the National Chung-Hsing University, Taiwan. Being the first new type of cell division to be unearthed in decades, asynthetic fission was observed when the team mapped a growing zebrafish embryo's skin cells in real time, some of which appeared to divide without replicating their DNA. The topic goes on - but this should at least indicate how important zebrafish can be for us.

New Models
In biochemistry, there are probably thousands of model organisms currently being explored around the world. While I've mentioned some of the most common of these, many others have seen the light of publications plenty of times before. The African clawed frog (Xenopus laevis) is one such organism, winning a number of Nobels for research behind its egg cells (which is more practical than it sounds). Nowadays, it is being replaced by the Western clawed frog (Xenopus tropicalis) - a smaller, more genetically simple kind of amphibian.
With the complex neurology of octopuses, scientists in Masachusetts have also invested much time in finding methods to domesticate them in the lab. Their efforts seem to bear fruit - though the ethics of confining intelligent beings like this are a topic in and of themselves. Fortunately, the majority of other model organisms are not so clever. I've not even mentioned plants, in this case, so it is safe to say that there is much more to model organisms than what we've seen. So, to answer the post's title, which is the best model organism?
I'm not sure - but it definitely depends on the type of research you're undertaking!
References
- Hedges, S. (2002). The origin and evolution of model organisms. Nature Reviews Genetics 3:838–849. Retrieved from https://doi.org/10.1038/nrg929
- Liu, W., Li, L., Ye, H., Chen, H., Shen, W., Zhong, Y., Tian, T., & He, H. (2017). From Saccharomyces cerevisiae to human: The important gene co-expression modules. Biomedical Reports 7(2):153–158. Retrieved from https://doi.org/10.3892/br.2017.941
- Hussain, N., Jenna, S., Glogauer, M. et al (2001). Endocytic protein intersectin-l regulates actin assembly via Cdc42 and N-WASP. Nature Cell Biology 3:927–932. Retrieved from https://doi.org/10.1038/ncb1001-927
- Farri, S. M. (2020). The rise to dominance of genetic model organisms and the decline of curiosity-driven organismal research. PLOS ONE 15(12):e0243088. Retrieved from https://doi.org/10.1371/journal.pone.0243088
- Chan, K. Y., Yan, C. C. S., Roan, H. Y. et al (2022). Skin cells undergo asynthetic fission to expand body surfaces in zebrafish. Nature. Retrieved from https://doi.org/10.1038/s41586-022-04641-0
- Grearson, A. G. et al (2021). The Lesser Pacific Striped Octopus, Octopus chierchiae: An Emerging Laboratory Model. Frontiers in Marine Science. Retrieved from https://doi.org/10.3389/fmars.2021.753483
